Heme-Associated Biofilm Remodeling in Mycobacterium abscessus
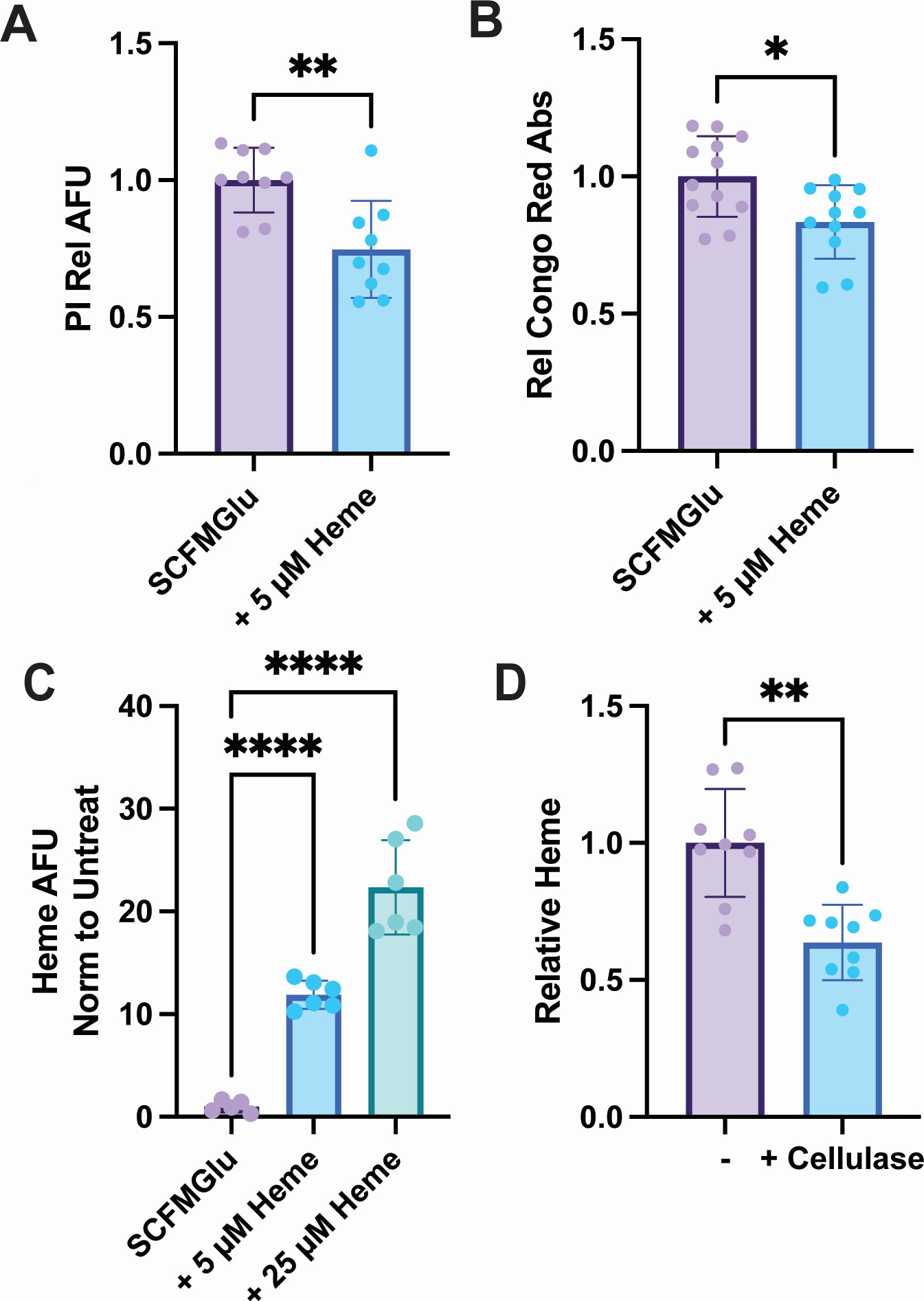
This client publication is relevant to teams studying how host-associated factors reshape bacterial physiology during chronic infection. In this study, extracellular heme altered the growth pattern and biofilm behavior of Mycobacterium abscessus, a pathogen closely associated with persistent infection and reduced antibiotic susceptibility.
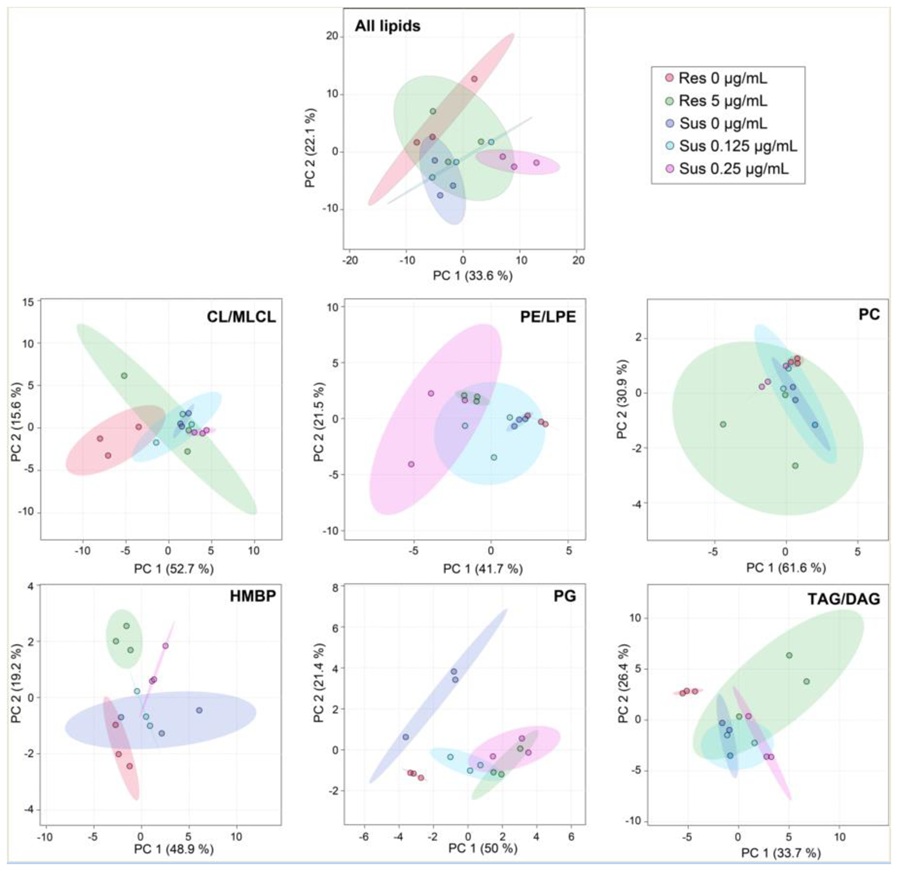
The study examined how exogenous heme influenced growth behavior, biofilm formation, and broader molecular responses in M. abscessus. The published work reports proteomics and metabolomics evidence, making this a strong case for biofilm- and membrane-associated molecular profiling, rather than a strictly lipidomics-only project.
The authors found that extracellular heme changed biofilm formation and broader metabolic behavior in M. abscessus. For service-page readers, the value of this case is that it shows how molecular profiling can help explain bacterial adaptation in persistent, barrier-associated states that are difficult to characterize with phenotype assays alone.
- Relevant to biofilm-associated infection research
- Useful for studies of persistence and reduced drug susceptibility
- Supports omics-guided interpretation of bacterial adaptation under host-like stress