Host Glycosphingolipids Essential for Viral Membrane Fusion
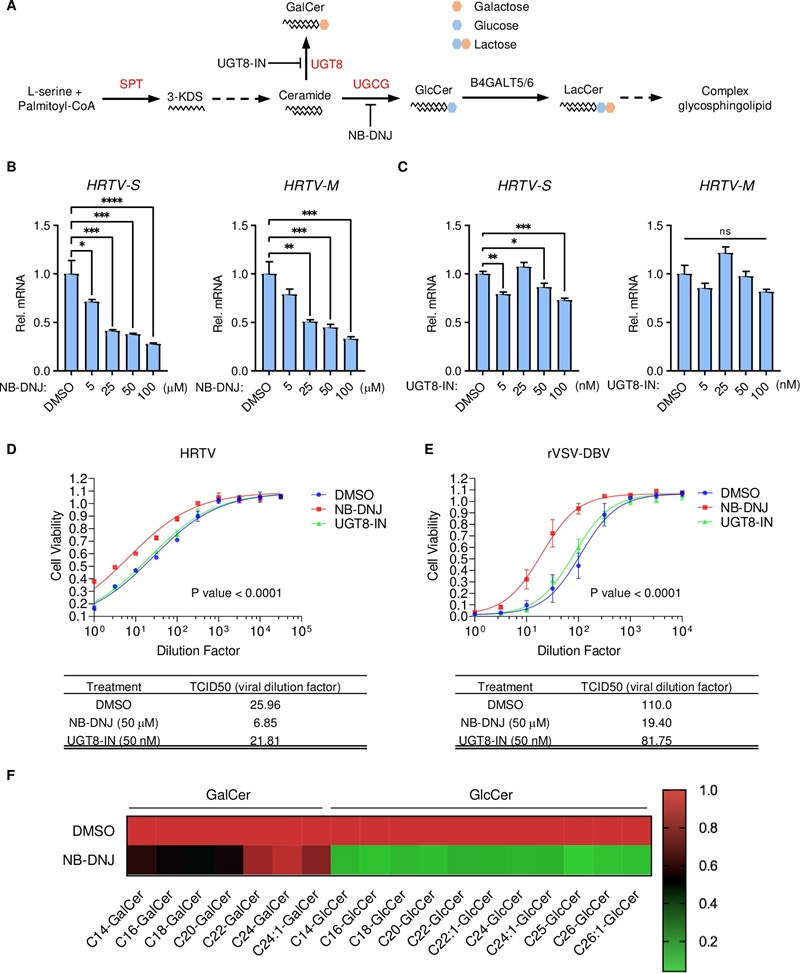
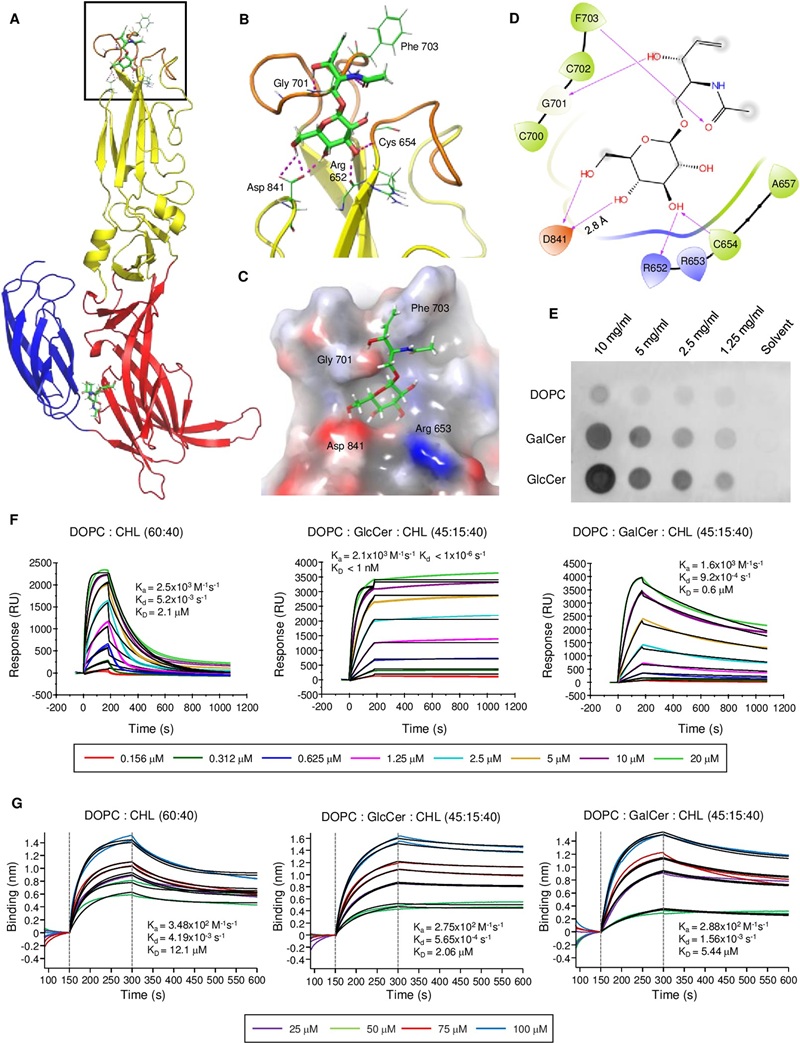
To understand how enveloped bandaviruses breach host cells, researchers utilized targeted host-pathogen lipidomics to quantify intracellular GalCer and GlcCer levels in 293T cells following pharmacological inhibition. This was coupled with Surface Plasmon Resonance (SPR) assays to evaluate HRTV Gc binding kinetics to specifically formulated liposomes.
The study directly proved that host glucosylceramide (GlcCer) is structurally essential for bandavirus glycoprotein-mediated membrane fusion. Depleting host GlcCer shifted lipid availability and effectively abolished viral entry, demonstrating a clear, targetable lipid-dependent infection mechanism.
Discovery Profiling → Targeted Sphingolipid Validation