Uncovering Biochemical Drivers of Nematode Resistance
Nematodes hijack host root tissues to form feeding sites. The client needed to prove that a specific resistance gene (*Rhg1*) actively alters the host's hormonal and lipid defense signaling. Measuring these transient, ultra-low-abundance defense molecules in infected roots requires extreme sensitivity.
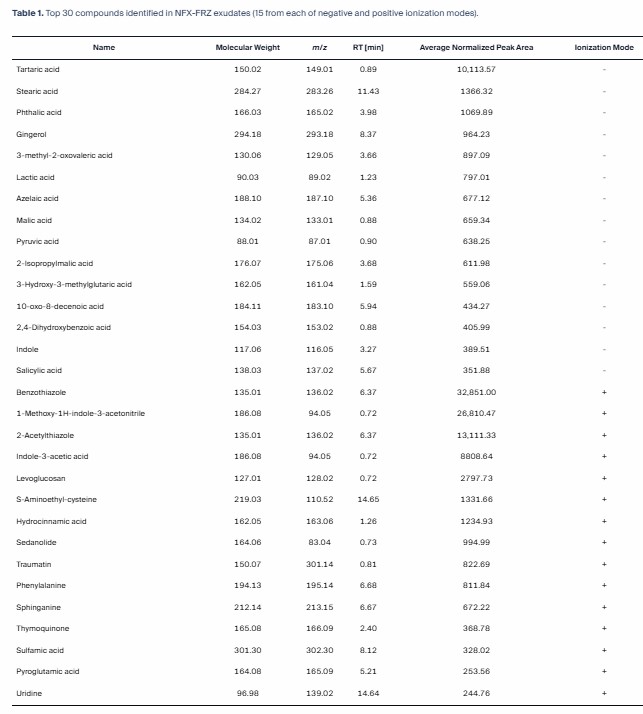
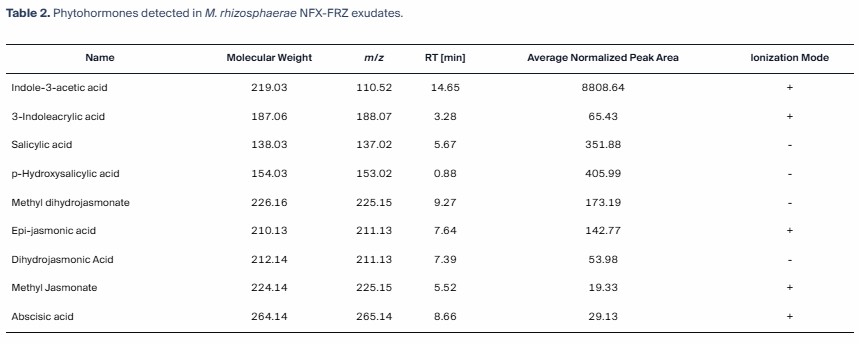
We deployed a highly sensitive Targeted LC-MS/MS workflow (Bundle B) to quantify defense-related phytohormones and lipid mediators. Our platform successfully tracked dynamic bursts of these signaling molecules across a precise infection time-course without matrix interference.
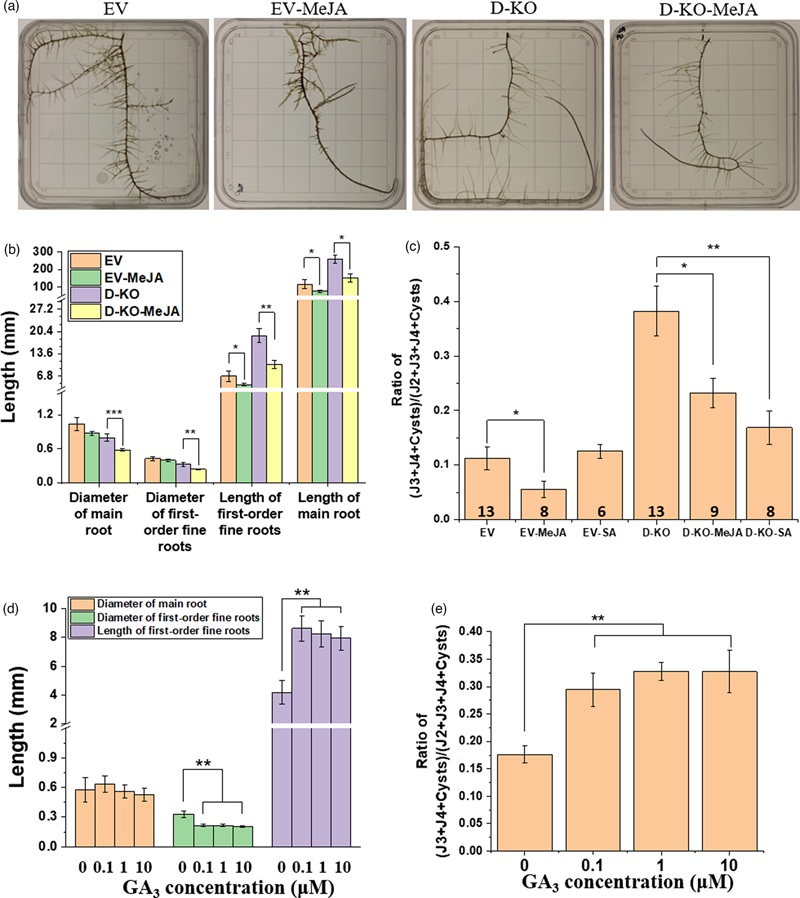
- Mechanism Defined: Revealed that *Rhg1* interacts with DELLA proteins to reprogram hormone pathways, restricting nematode feeding.
- Breeding Marker: Provided a definitive biochemical marker to guide the selection of resistant crop cultivars.