Discovery-Stage Profiling for a New Microalgal Production Chassis
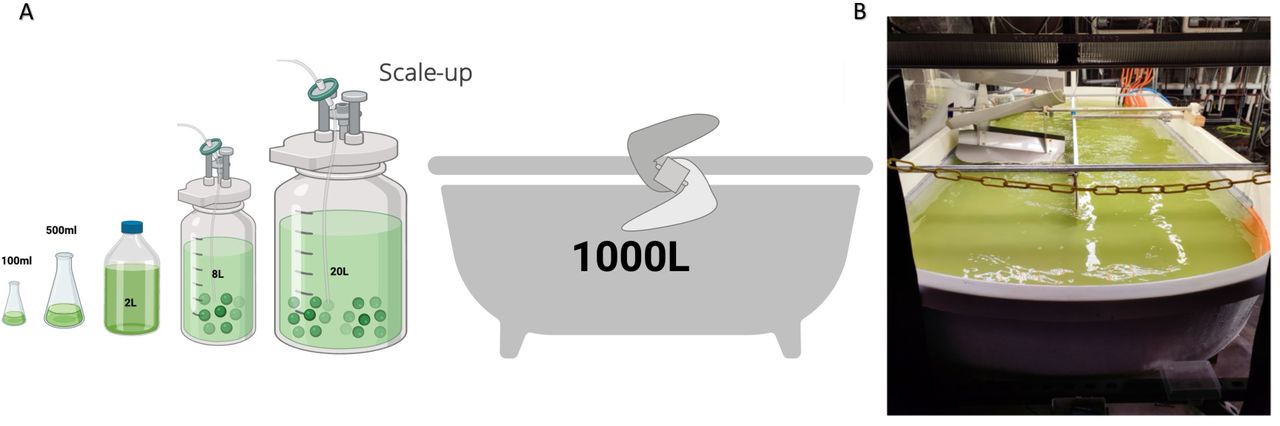
Early-stage microbial cell factory programs often begin with a basic but important question: is this strain worth advancing into deeper development? This client publication focused on Micractinium rhizosphaerae NFX-FRZ, a rhizosphere-associated microalga characterized for its biological and chemical output profile.
For customers evaluating a new microbial or algal chassis, discovery-stage profiling can help determine whether the organism shows enough molecular complexity and application potential to justify strain optimization, pathway follow-up, or bioprocess development. This is particularly relevant when target analytes have not yet been fully defined.
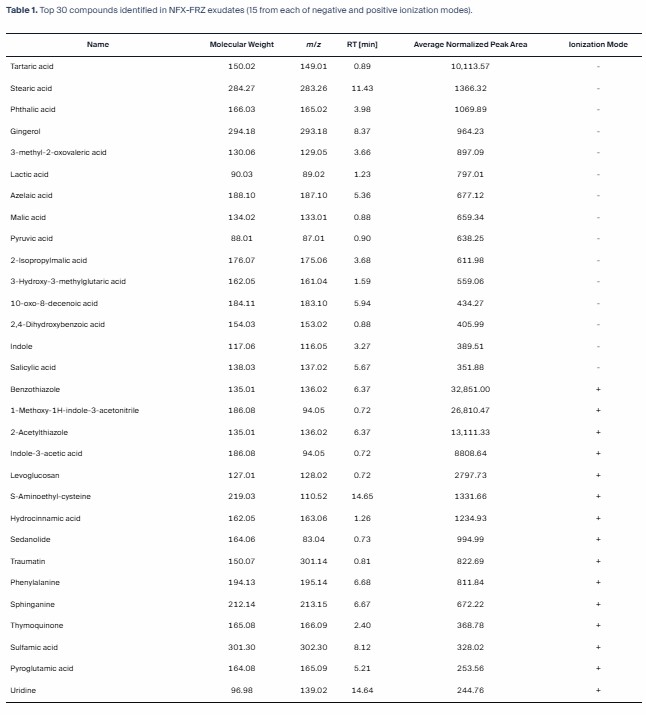
The article reports broad metabolomic characterization of NFX-FRZ exudates and shows that the strain produces multiple biologically relevant compounds, including organic acids and phytohormone-related molecules. In a service-page context, this case illustrates the value of broad profiling when a team needs an evidence-based starting point for chassis evaluation rather than a single predefined assay.
- Evaluating a newly isolated microalgal or microbial strain
- Comparing non-model strains before pathway engineering
- Building a molecular evidence base for downstream production development