Feedstock-Linked Fermentation Inhibition Diagnosis in Yeast Bioethanol Production
This study is highly relevant to teams investigating why lignocellulosic feedstocks perform inconsistently during yeast fermentation, especially when process failure may originate upstream from biomass quality rather than downstream reactor settings. It is best presented as a fermentation troubleshooting case, not as a pure dynamic lipid remodeling study.
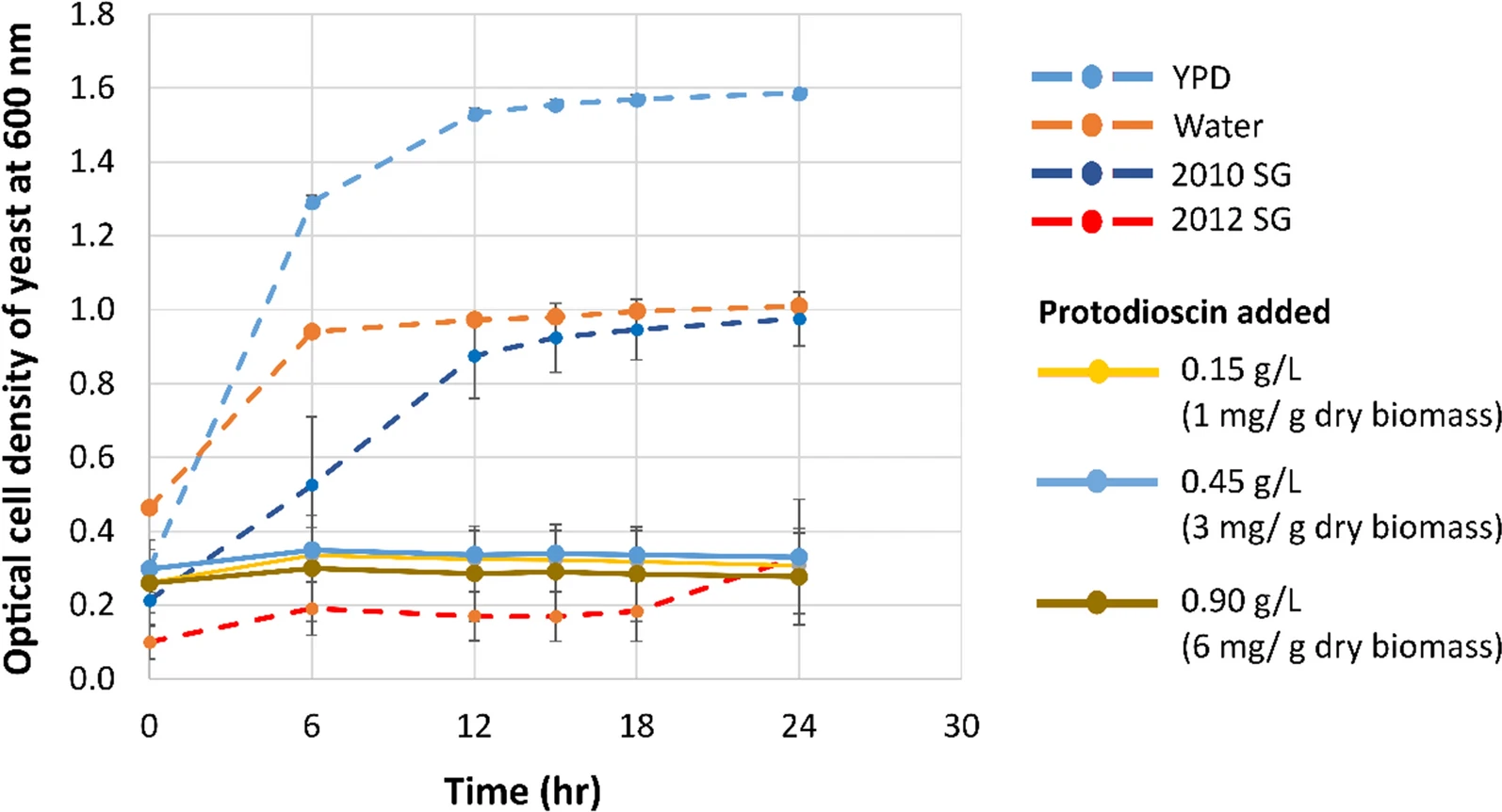
The authors compared control-year and drought-year switchgrass, combined extraction and pretreatment workflows, and evaluated fermentation performance alongside LC-MS-based characterization of inhibitory compounds enriched in the hydrolysates. The study then used protodioscin as a commercially available saponin surrogate to test whether added saponin burden could reproduce yeast growth inhibition.
The paper linked drought-associated switchgrass chemistry to reduced yeast growth and fermentation performance, and identified water-soluble saponin enrichment as a plausible inhibitory factor. For external-facing content, the value of this case lies in showing how complex fermentation underperformance can be traced back to chemically defined feedstock-derived inhibitors rather than treated as a generic bioreactor issue.
For bioprocess groups screening variable lignocellulosic inputs, this case supports a more defensible workflow for inhibitor diagnosis, raw-material triage, and pre-fermentation risk assessment. It also demonstrates capability in handling complex biomass-derived matrices and converting broad process symptoms into molecule-level explanations.