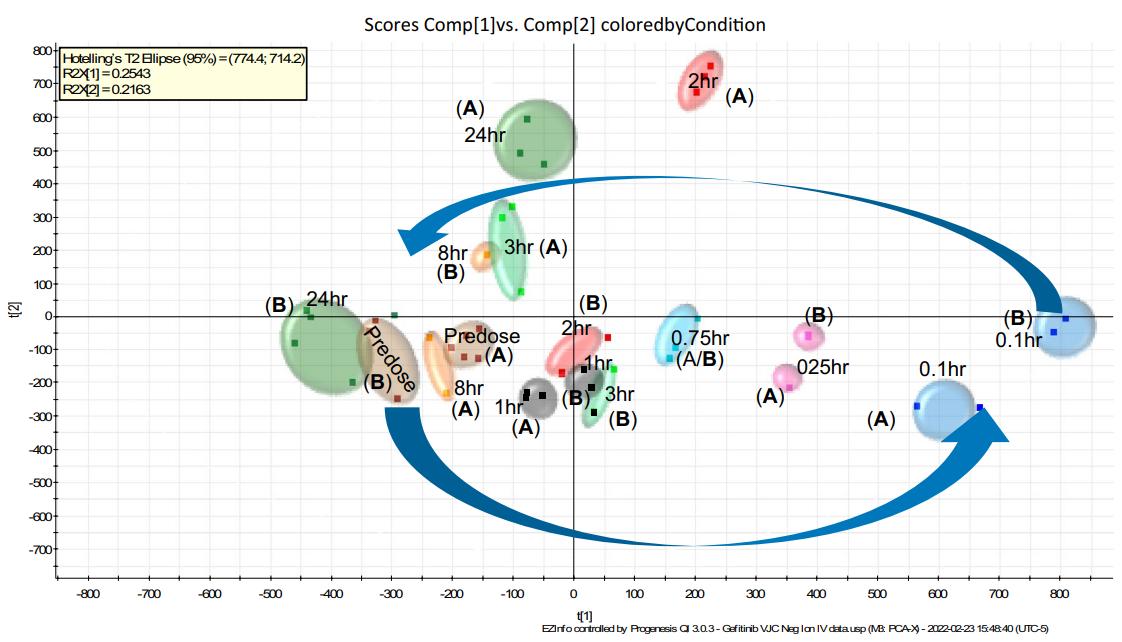
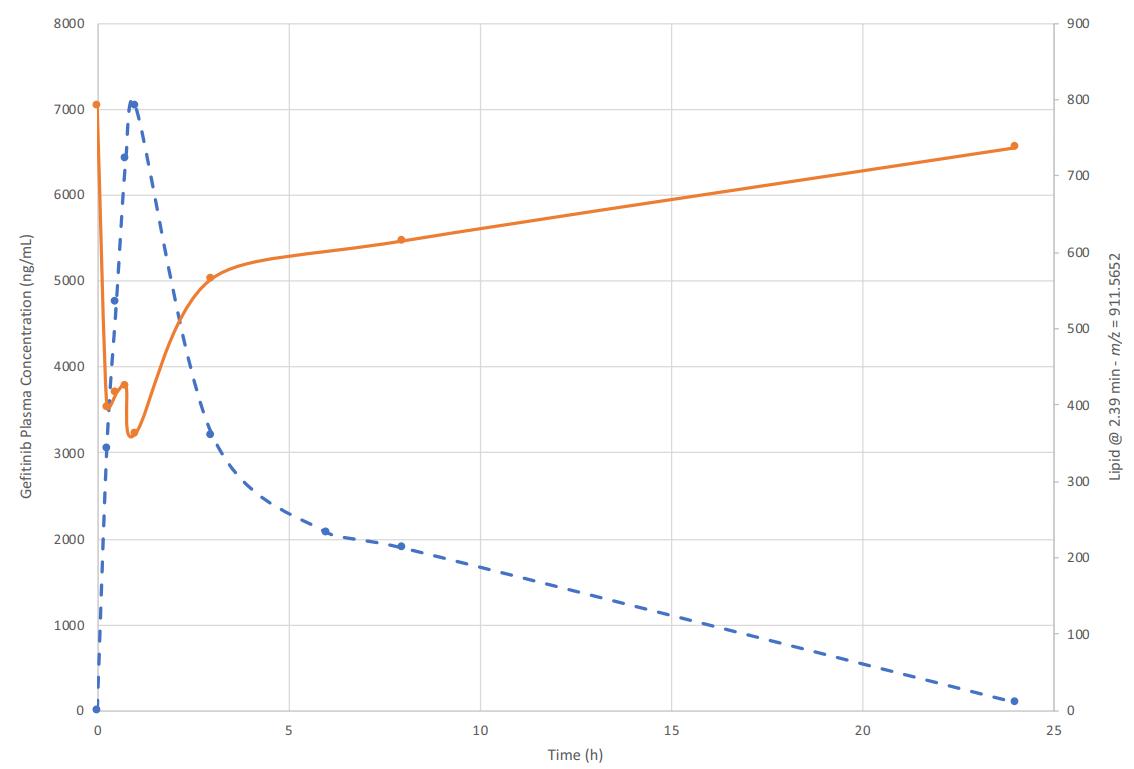
Situation: Plasma readouts are subtle or inconsistent across post-dose timepoints; treated vs control separation is near baseline variability, or directionality flips across adjacent timepoints.
Goal: Identify exposure-aligned lipid pathways and convert them into quant-stable PD markers suitable for confirmation and longitudinal tracking.
Recommended Path: Bundle A → Bundle B
Recommended Services:
What You Will Get: Exposure-aligned lipid signature candidates plus a clear conversion path into targeted panels that are more stable across time, dose, and batches.
Situation: Lead optimization requires repeat sampling across cohorts and dose levels; plasma/serum must support longitudinal PD/PK tracking.
Goal: Establish a minimally invasive lipid biomarker set and translate it into targeted panels for repeatable monitoring (RUO).
Recommended Path: Bundle A → Bundle B
Recommended Services:
What You Will Get: A plasma/serum PD/PK marker strategy built around targeted panels that are easier to replicate across cohorts and timepoints.
Situation: PD signals don't replicate across batches/sites; effect sizes don't match exposure; likely pathway-specific markers are being diluted by broad profiling or matrix/panel mismatch.
Goal: Rebuild a reproducible PD readout with panel-first endpoints that survive site/batch variation.
Recommended Path: Bundle B (panel-first) → optional Bundle A (context)
Recommended Services:
What You Will Get: A narrowed, reproducibility-first PD panel set with pathway specificity (sphingolipid/sterol modules) that is more stable across sites and batches.
Situation: Evidence suggests membrane remodeling (PC/PE/PI, DAG/TAG shifts, ratios), but discovery-level changes aren't robust enough for lead ranking across dose/time/batch.
Goal: Quantify remodeling endpoints with higher confidence and mechanistic specificity to support PD interpretation and optimization decisions.
Recommended Path: Bundle B
Recommended Services:
What You Will Get: Quant-stable membrane remodeling readouts (phospholipid + neutral lipid modules) plus energy/β-oxidation context when acylcarnitines are relevant.
Situation: Safety pharmacology or efficacy phenotype suggests inflammation signaling; you need mediator-layer PD rather than bulk membrane lipids alone.
Goal: Quantify mediators and connect changes to COX/LOX/CYP pathway branches to support PD biology and risk interpretation.
Recommended Path: Bundle B (mediators)
Recommended Services:
What You Will Get: A mediator-centric PD package (oxylipins/eicosanoids/PGs) that is closer to inflammatory mechanism than bulk lipid classes, with optional upstream lipid drivers.
Situation: Plasma biomarkers move with treatment, but efficacy doesn't correlate; tissue localization, compartment effects, or mitochondrial involvement is suspected.
Goal: Anchor lipid changes to the site of action and identify which lipid modules best reflect pharmacology in tissue.
Recommended Path: Bundle A (tissue bridge) → Bundle B
Recommended Services:
What You Will Get: Tissue-anchored PD interpretation with mitochondrial/membrane modules that better reflect target engagement at the site of action.
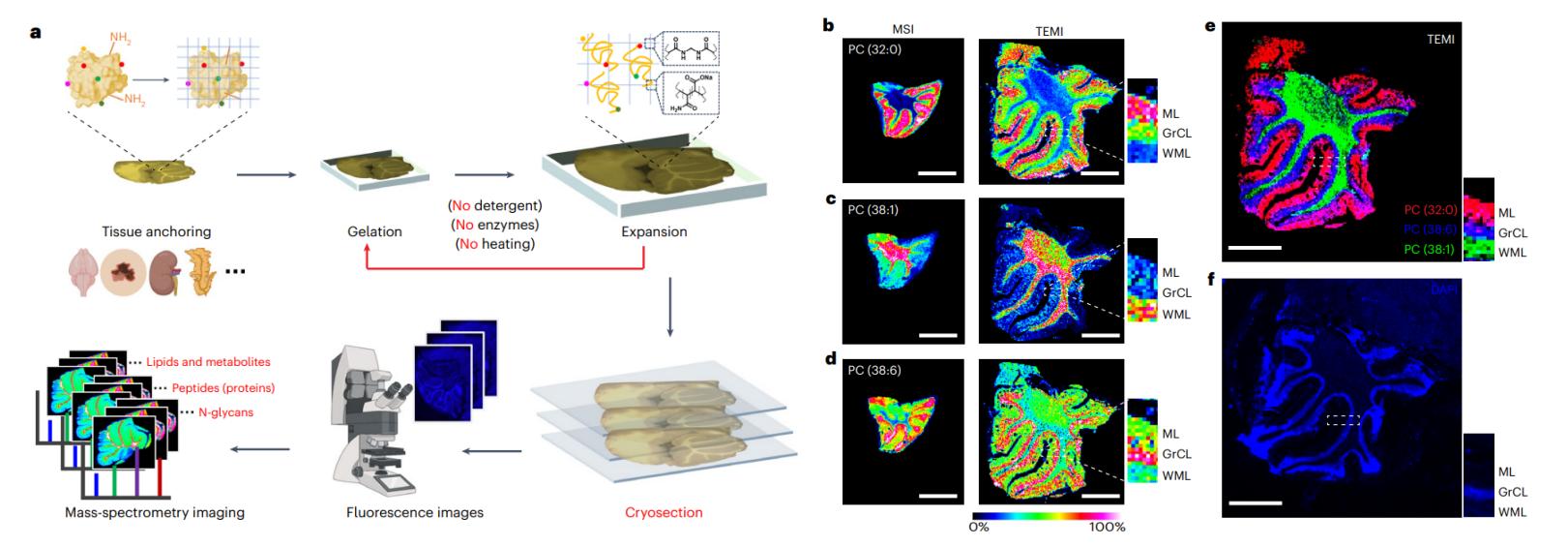
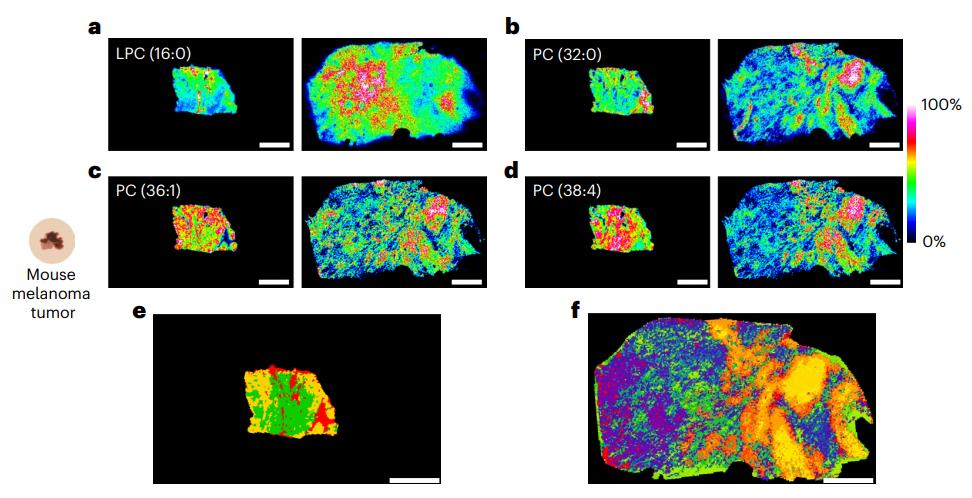
Situation: Heterogeneous tissues dilute localized lipid responses; bulk tissue appears "flat" despite suspected regional pharmacology.
Goal: Visualize spatial lipid changes and quantify region-level differences that explain responder/non-responder patterns or microenvironment effects.
Recommended Path: Bundle C (imaging)
Recommended Services:
What You Will Get: Spatial lipid maps plus region-stratified marker candidates that explain localized PD even when bulk averages fail.
Situation: Steady-state lipid shifts have multiple interpretations; decision-makers require stronger causal evidence (synthesis vs turnover) for MoA/target engagement.
Goal: Provide flux-informed evidence that clarifies whether exposure changes lipid production, remodeling, or turnover.
Recommended Path: Bundle C (flux) → Bundle B
Recommended Services:
What You Will Get: Flux-backed pathway interpretation that separates synthesis vs turnover, supporting stronger PD/PK biomarker decisions at program inflection points.