Microbiome Diversity and Metabolic Interplay in the Zebrafish Intestine
The zebrafish is a premier aquatic model, but characterizing the minute metabolic environment of the dissected larval intestinal tract poses severe sensitivity challenges. The client needed to prove that specific pathogen secretion systems disrupt the microbial ecosystem, which in turn feeds back into the host's metabolic status.
We deployed a micro-volume Untargeted Metabolomics and Lipidomics workflow. By leveraging the extreme sensitivity of the Orbitrap mass spectrometry platform, we successfully captured the metabolic signals from dissected zebrafish larval intestines. This was integrated with microbial structural analysis to correlate diversity shifts with metabolic disruption.
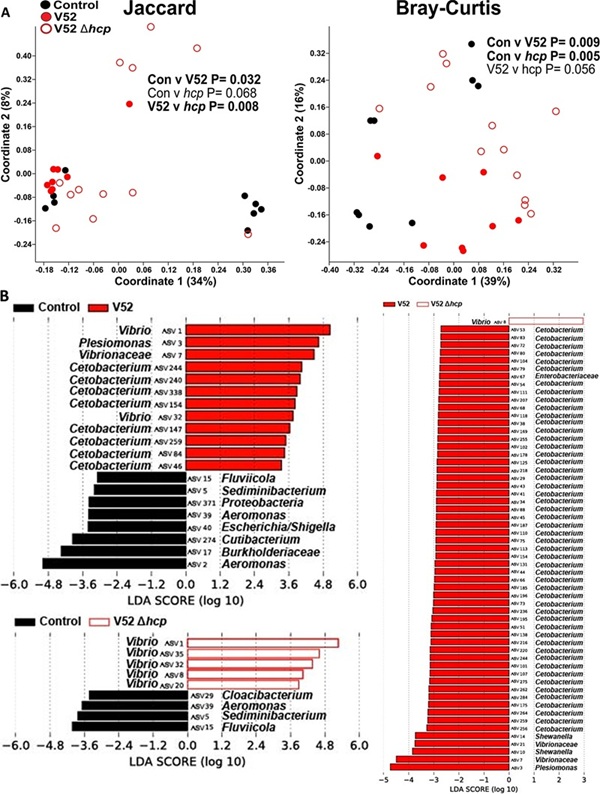
The analysis demonstrated clear shifts in the β-diversity of the zebrafish intestinal microbiome following infection using PCoA plots. These plots provided the essential structural evidence that the pathogen significantly alters the microbial community. This framework empowers researchers to link microbiome structural failure to the subsequent collapse of the host’s gut lipid barrier.