Untargeted betaine lipidomics profiling
Designed for broad DGTS/DGTA/DGCC coverage and discovery comparisons. This option emphasizes high-resolution MS and MS/MS-supported annotation.
Our services have earned the trust of companies, schools, and organizations globally, and we remain dedicated to maintaining that trust.
Betaine lipids are polar lipids with choline-like chemistry but no phosphate. In several biological systems, they are linked to membrane adaptation under nutrient shifts and stress response.
You typically analyze DGTS/DGTA/DGCC when you need to:
A key technical point matters for planning: DGTS and DGTA are structural isomers. They can produce similar MS/MS patterns, so identification confidence must be documented.
Choose a solution based on whether you need discovery coverage or quant tracking.
Untargeted betaine lipidomics profiling
Designed for broad DGTS/DGTA/DGCC coverage and discovery comparisons. This option emphasizes high-resolution MS and MS/MS-supported annotation.
Targeted DGTS/DGTA/DGCC panel quantification
Designed for consistent tracking across conditions or batches. This option emphasizes triple quadrupole MRM-based quant workflows and QC reporting.
MS/MS evidence pack for higher-confidence IDs
Designed for projects that need an audit trail for key calls. You receive representative spectra and annotation notes for selected lipids.
Membrane remodeling add-on
Designed to contextualize betaine lipids alongside major polar lipid classes. This supports interpretation of substitution patterns.
| Class / Category | What we detect | Typical reportable species range | Representative examples | Notes on confidence |
|---|---|---|---|---|
| DGTS (Diacylglyceryl trimethylhomoserine) | Core betaine lipid class in many algae/microbes/plants | C28–C44 : 0–11 | 30:0, 32:0, 32:1, 34:2, 36:4, 38:6, 40:6, 42:10 | Isomer-aware where DGTS overlaps with DGTA; MS/MS evidence provided when requested |
| DGTA (Diacylglyceryl hydroxymethyltrimethyl-β-alanine) | DGTS structural isomer; often matrix- and taxa-dependent | C30–C44 : 0–10 (project-dependent) | 32:1, 34:2, 36:4, 38:6 | Requires careful LC separation + MS/MS; ambiguity is flagged, not forced |
| DGCC (Diacylglyceryl carboxyhydroxymethylcholine) | Commonly seen in marine phytoplankton lipidomes | C32–C44 : 0–10 | 32:x, 34:x, 36:x, 38:x, 40:x, 42:x, 44:x (x = unsaturation) | Often reported as sum compositions; MS/MS used to support class confirmation |
| MGTS (Lyso / monoacyl-GTS) | Turnover/remodeling intermediate | C14–C22 : 0–6 | 16:0, 16:1, 18:1, 18:2, 18:3, 20:5 | Reported when present above detection thresholds; useful for remodeling context |
| MGCC (Lyso / monoacyl-GCC) | DGCC-related lyso form | C14–C22 : 0–6 (matrix-dependent) | 16:x, 18:x, 20:x | Often lower abundance; detection depends on biomass type and extraction |
| Optional output | What it adds | Typical use cases |
|---|---|---|
| Fatty-acyl motif summaries (e.g., EPA/DHA/ARA-containing species) | Adds composition-level interpretation beyond C:D | Membrane remodeling under stress; strain comparisons; marine phytoplankton profiling |
| MS/MS evidence pack for selected species | Spectral confirmation and annotation transparency | DGTS vs DGTA differentiation; publication-grade figures |
We apply annotation rules that acknowledge isomer ambiguity when it exists. We do not force a single label without evidence.
We provide LC separation context and MS/MS support for selected targets. This reduces misassignment risk in DGTS/PC-dense regions.
For targeted studies, we report calibration and QC summaries alongside results. Common acceptance targets are used where appropriate, such as:
When relevant, we include DGTS/PC ratio and PC-to-DGTS substitution ratio outputs to support membrane remodeling interpretation.
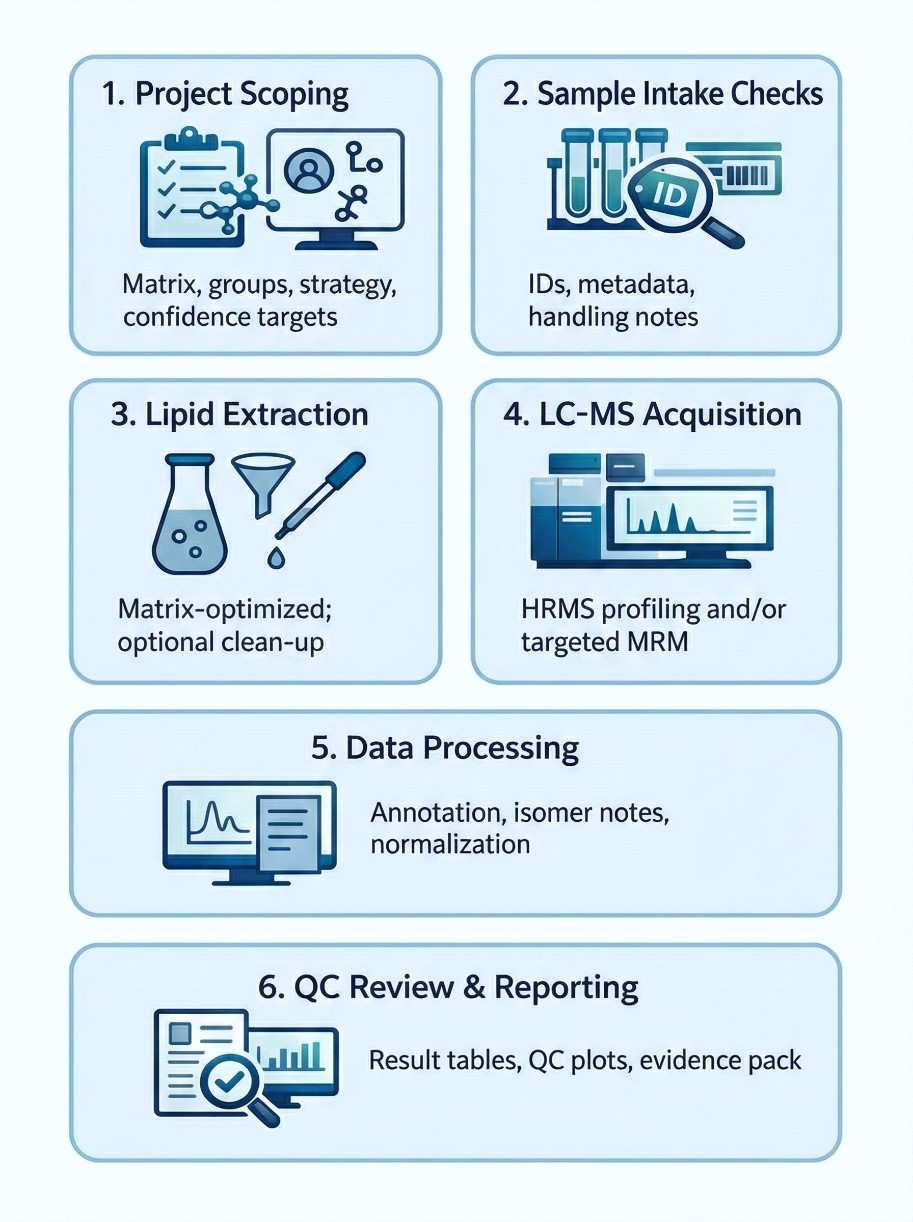
We combine UHPLC separation with high-resolution MS for profiling/confirmation and triple quadrupole LC–MS/MS for targeted quantification. Platform choice follows your decision need.
| Aspect | Profiling & confirmation | Targeted quantification |
|---|---|---|
| MS platform | Orbitrap Exploris 480 | Agilent 6495C |
| MS approach | HRMS with MS/MS | Triple quadrupole MRM |
| LC separation | RP-UHPLC (C18) and/or HILIC | RP-UHPLC (commonly) and/or HILIC |
| Ion source | ESI; polarity selected by class and matrix | ESI; polarity selected by class and matrix |
| Key parameters documented | Resolving power, mass accuracy strategy, MS/MS settings | Transitions, collision energy, dwell time |
| Output style | Relative abundance matrices; optional MS/MS evidence | Concentrations or QC-anchored normalized abundances |
| Evidence level | Exact-mass + MS/MS-supported annotation | MRM transitions with QC summaries |
| QC context | Blanks, internal standard behavior, run-level checks | Blanks, internal standard behavior, run-level checks |

Agilent 6495C Triple Quadrupole (Figure from Agilent)

Agilent 1260 Infinity II HPLC (Fig from Agilent)

Orbitrap Exploris 480 (Figure from Thermo)
Included with every betaine lipidomics project.
Added when deeper comparison or interpretation is required.
All deliverables are provided in formats compatible with common downstream workflows.
Explore our Lipidomics Solutions brochure to learn more about our comprehensive lipidomics analysis platform.

Membrane remodeling studies
Quantify DGTS/DGTA/DGCC shifts to map lipid headgroup substitution.

Phosphate limitation & nutrient stress
Track phosphorus-free lipid adoption when PC-based signals stall.

Algal strain screening
Compare betaine lipid fingerprints for selection, engineering, and productivity studies.

Plant & microbial stress biology
Link betaine lipid patterns to adaptation phenotypes across complex matrices.

Marine & environmental lipidomics
Profile DGCC-rich signatures in biomass and community samples.

Multi-omics mechanism building
Integrate betaine lipid readouts with transcriptomics/proteomics for pathways.
| Sample type | Suggested amount | Recommended format | Handling | Notes (keep it short) |
|---|---|---|---|---|
| Algal biomass / slurry | 20–100 mg dry biomass or 0.5–2 mL dense slurry | Wet slurry, paste, or pellet | Keep cold; sealed; protect from light | Species/strain; condition (e.g., P limitation); biomass basis (dry/wet) |
| Microbial cultures (bacteria/yeast/fungi) | ≥1×10⁷ cells or 10–50 mg wet pellet | Cell pellet (preferred) | Keep cold; minimize residual medium | Strain; medium; stress/induction; wash notes (if any) |
| Plant tissue | 20–50 mg | Cryo-powder or small pieces | Keep cold; dry-sealed; protect from light | Tissue type; treatment; harvest metadata |
| Environmental biomass | 1–3 filters or 10–50 mg pellet | Filters or biomass pellet | Keep cold; sealed; protect from light | Filter type; volume processed; field blanks if available |
| Isolated membranes (optional) | 50–200 µg protein (or equivalent) | Membrane fraction pellet | Keep cold; minimal additives | Buffer composition; detergent presence (if any) |
| Client-prepared lipid extracts (optional) | Extract equivalent to ≥20 mg biomass/tissue | Dried extract or organic extract | Sealed; protect from light; minimal headspace | Solvent system; extraction method; internal standards (if used) |
| Plasma / serum (special-case) | 100–300 µL | Clarified plasma/serum | Keep cold; avoid hemolysis; protect from light | Dietary exposure / tracer studies only; fed/fasted note if relevant |
| Whole blood (generally not recommended) | ≥200 µL (only if required) | Whole blood aliquot | High matrix; avoid unless justified | Anticoagulant type; handling notes |
Note: If your sample is limited, we can prioritize a targeted panel or a reduced deliverable set to match feasibility.
Q: How do you distinguish between DGTS and DGTA isomers, given they have the same mass?
A: We separate DGTS and DGTA using UHPLC chromatography based on their interaction with the stationary phase. MS/MS fragmentation further confirms their identity, distinguishing DGTS (homoserine headgroup) from DGTA (hydroxymethyltrimethyl-β-alanine headgroup).
Q: Can this assay quantify betaine lipids in the presence of high Phosphatidylcholine (PC) levels?
A: Yes. Our method uses high-resolution MS (HRMS) or targeted MRM transitions to filter out PC interference, ensuring accurate quantification of betaine lipids even in PC-dominant samples.
Q: Is the data reported as absolute concentration or relative abundance?
A: We offer:
Q: Should I extract the lipids myself or send raw biomass?
A: We recommend sending raw biomass (frozen pellets or tissue) to ensure maximum recovery and avoid lipid degradation. Contact us for solvent compatibility if you need to send extracts.
Q: My samples are marine algae with high salt content. Will this affect the LC-MS analysis?
A: We include a desalination step and use chromatography to eliminate salt interference, maintaining high sensitivity in marine samples.
Q: Can you calculate the DGTS/PC ratio to assess phosphorus limitation?
A: Yes. We recommend adding our "Major Polar Lipids" module, which quantifies both DGTS and PC to calculate their ratio and assess phosphorus limitation.
Q: Do you report fatty acyl composition for each betaine lipid species?
A: Yes, for abundant species. We report specific fatty-acyl chains (e.g., DGTS 16:0/20:5) or sum compositions (e.g., DGTS 36:5) for lower-abundance species.

Services:
Online Inquiry
CONTACT US