PFAS-Driven Lipid Remodeling Linked to Early Immune Injury
Female NOD mice were exposed to perfluoroundecanoic acid (PFUnDA) during gestation, lactation, and early life. Serum samples from offspring were analyzed by comprehensive lipidomics, and the lipid data were evaluated against exposure level, insulitis grade, macrophage counts, and apoptotic markers in pancreatic islets.
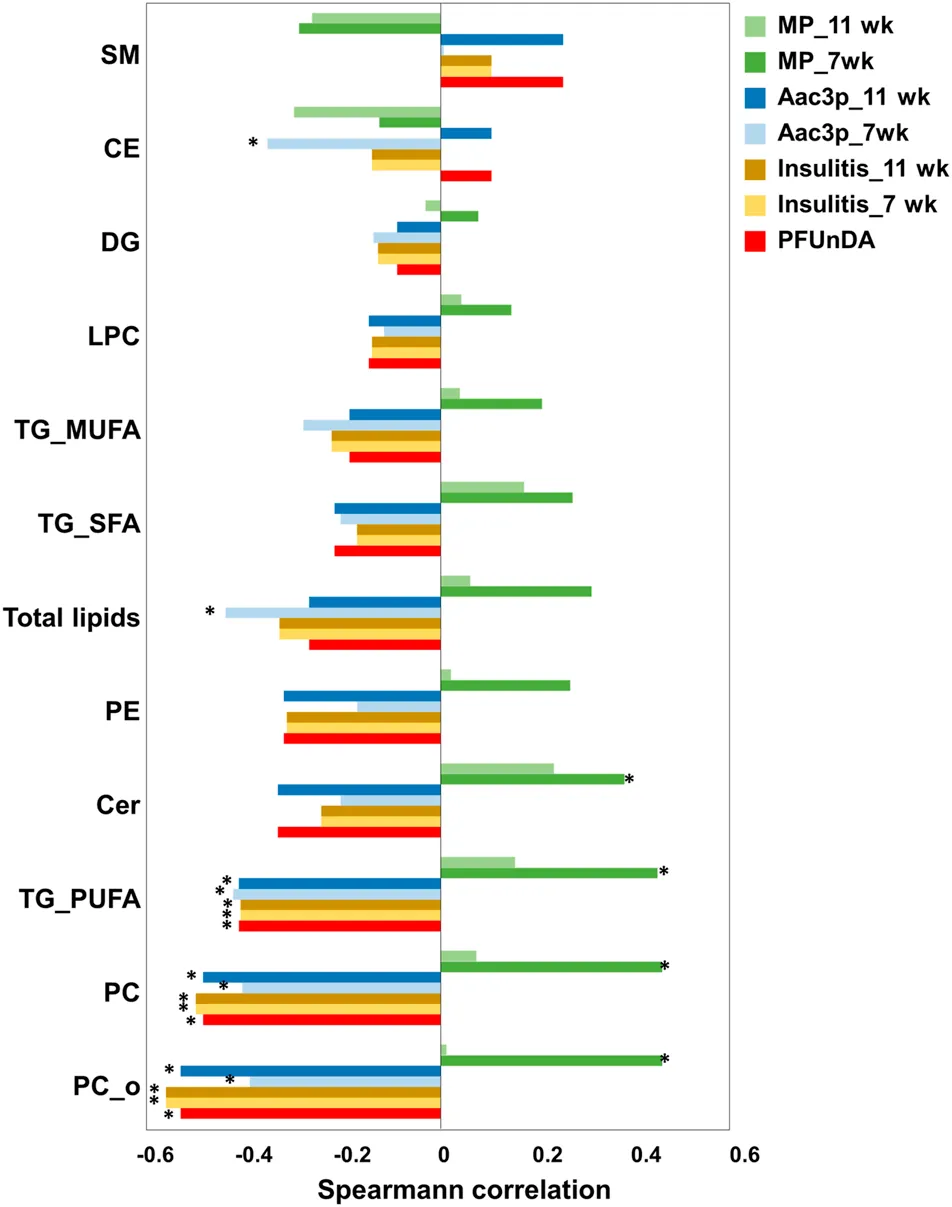
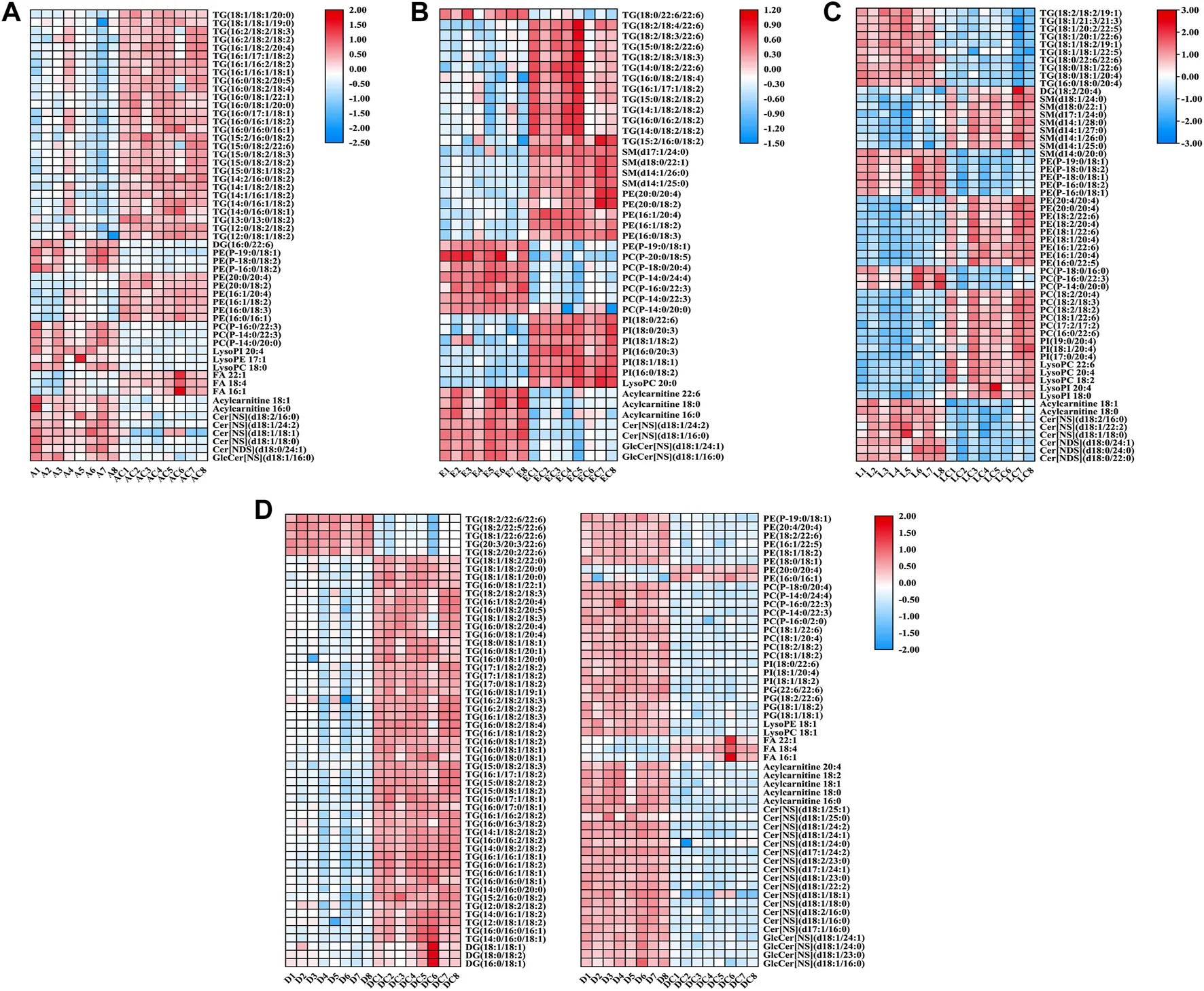
PFUnDA exposure produced dose-dependent lipidomic shifts, with reduced phosphatidylcholines, ether phosphatidylcholines, and PUFA-containing triacylglycerols. These lipid changes also tracked with insulitis-related endpoints, making the study a strong example of how mechanistic toxicology lipidomics can connect environmental exposure to pathway-level injury signals rather than phenotype alone.
Bundle A (Untargeted Lipidomics Profiling) → Bundle B (Targeted Validation of phospholipids/sphingolipids) → Bioinformatic correlation analysis