Targeted Sphingolipid Quantitation for Membrane-Fusion MoA Support
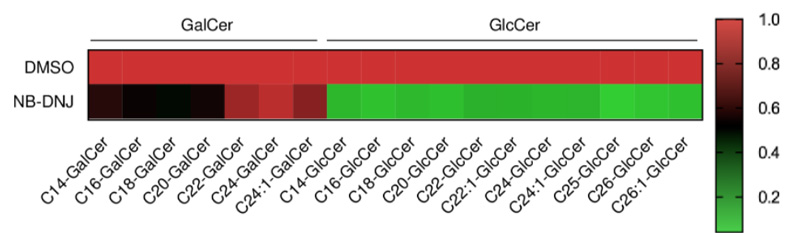
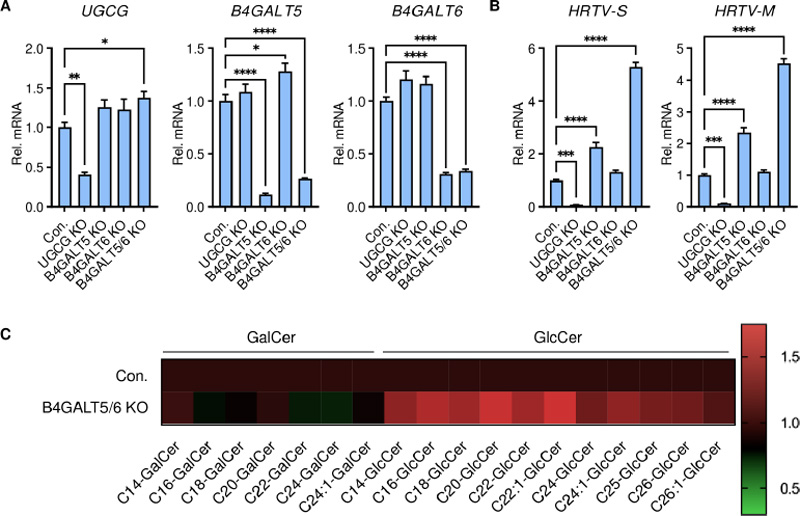
Precisely track structurally similar glycosphingolipids (e.g., GlcCer vs GalCer) and determine whether the pathway signal reflects causal MoA involvement (entry/fusion) rather than downstream stress effects—something that broad “omics” layers often fail to resolve.
We recommend a Bundle B validation path centered on targeted lipidomics (LC–MS/MS, MRM/PRM) for sphingolipid-class quantitation, with a panel designed around MoA hypotheses (glycosphingolipid biosynthesis / remodeling). This enables clear pathway engagement readouts across inhibitor treatment, genetic perturbation, or time-course designs.
- MoA anchoring: Quantitative sphingolipid readouts provide a direct biochemical anchor for mechanism hypotheses when membrane biology is central.
- Cleaner target deconvolution: Helps prioritize upstream enzymes/transporters as candidate targets and guides follow-up validation experiments.
- Formulation relevance (optional extension): The same lipid classes are also relevant to vesicle membranes and lipid-based delivery systems, supporting cross-program learning.