Developmental Lipidomics in Sesame Seeds
This study closely matches projects that need to identify the decisive developmental window for oil accumulation and connect lipid changes to candidate biosynthetic genes. It is particularly useful for teams planning stage-based sampling, multi-omics integration, or pathway validation.
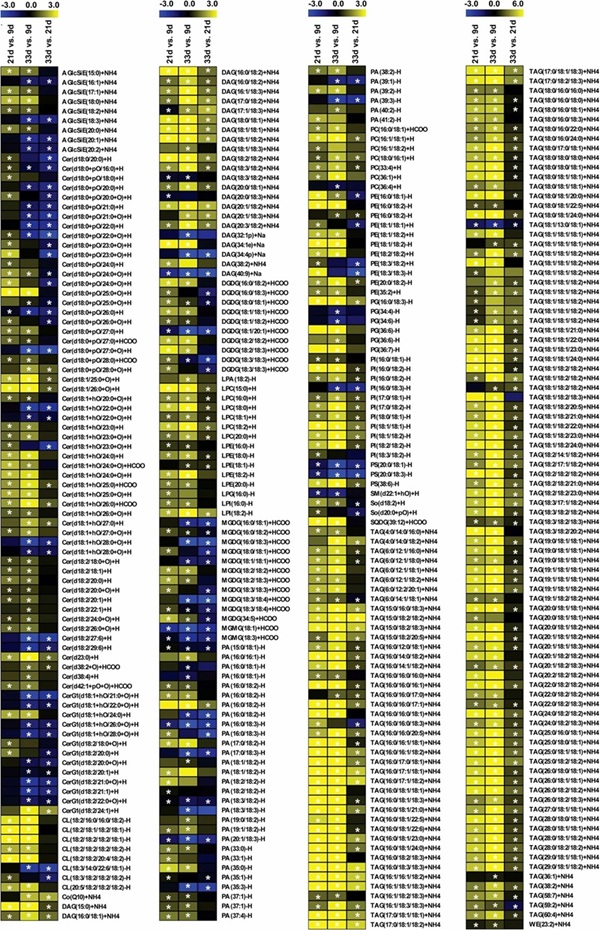
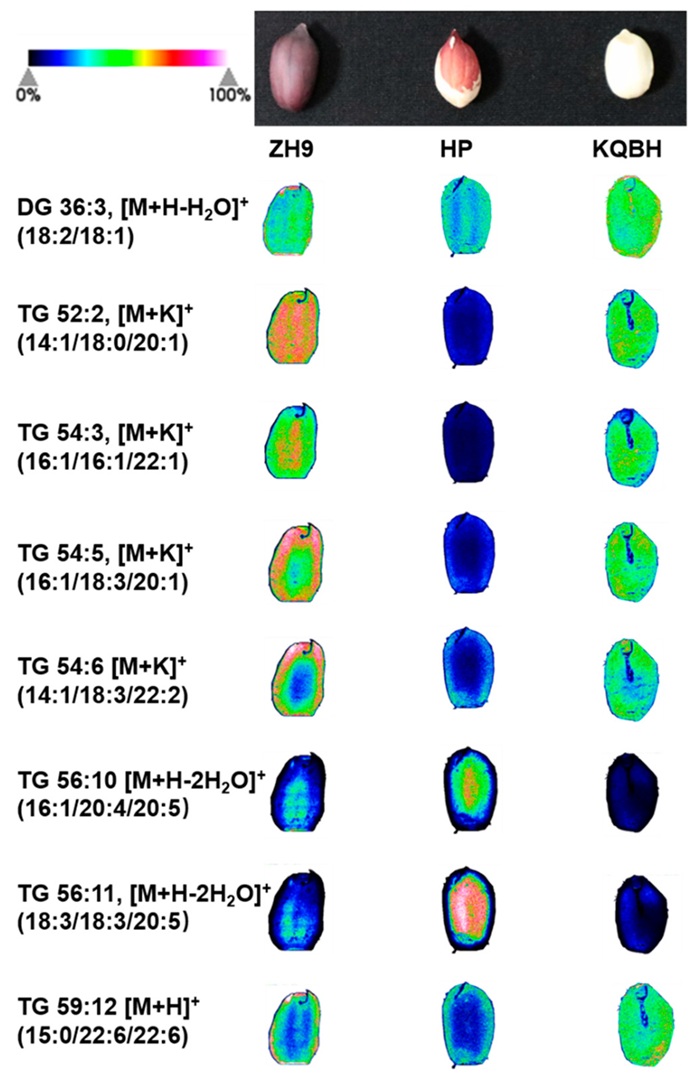
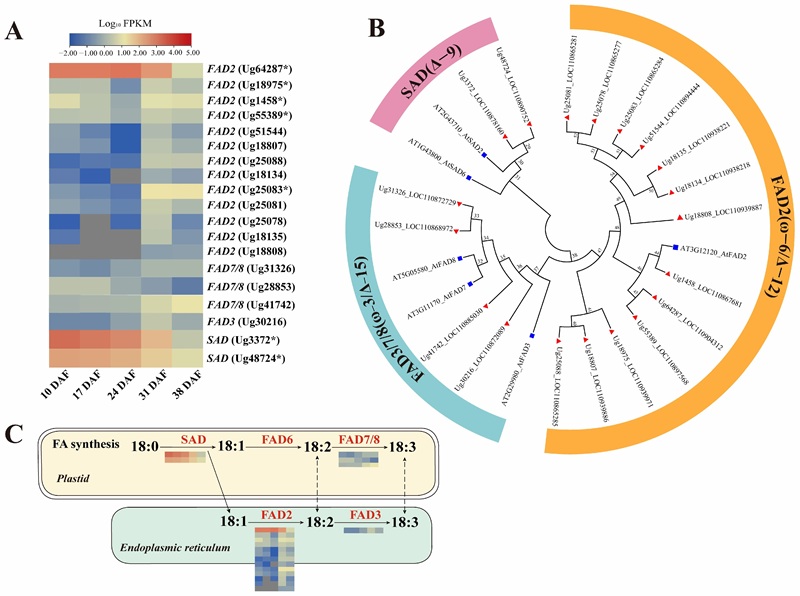
Sesame seeds sampled at 9, 21, and 33 days after flowering were analyzed by LC-MS lipidomics, GC-MS fatty acid analysis, and transcriptomics. The study reported 481 lipids and showed that most fatty acids and storage-related lipids increased most strongly at the middle and late stages, with accompanying expression changes in genes including ACCase, FAD2, DGAT, GPAT, LPAT, PAP, and WRI1-like regulators.