Situation
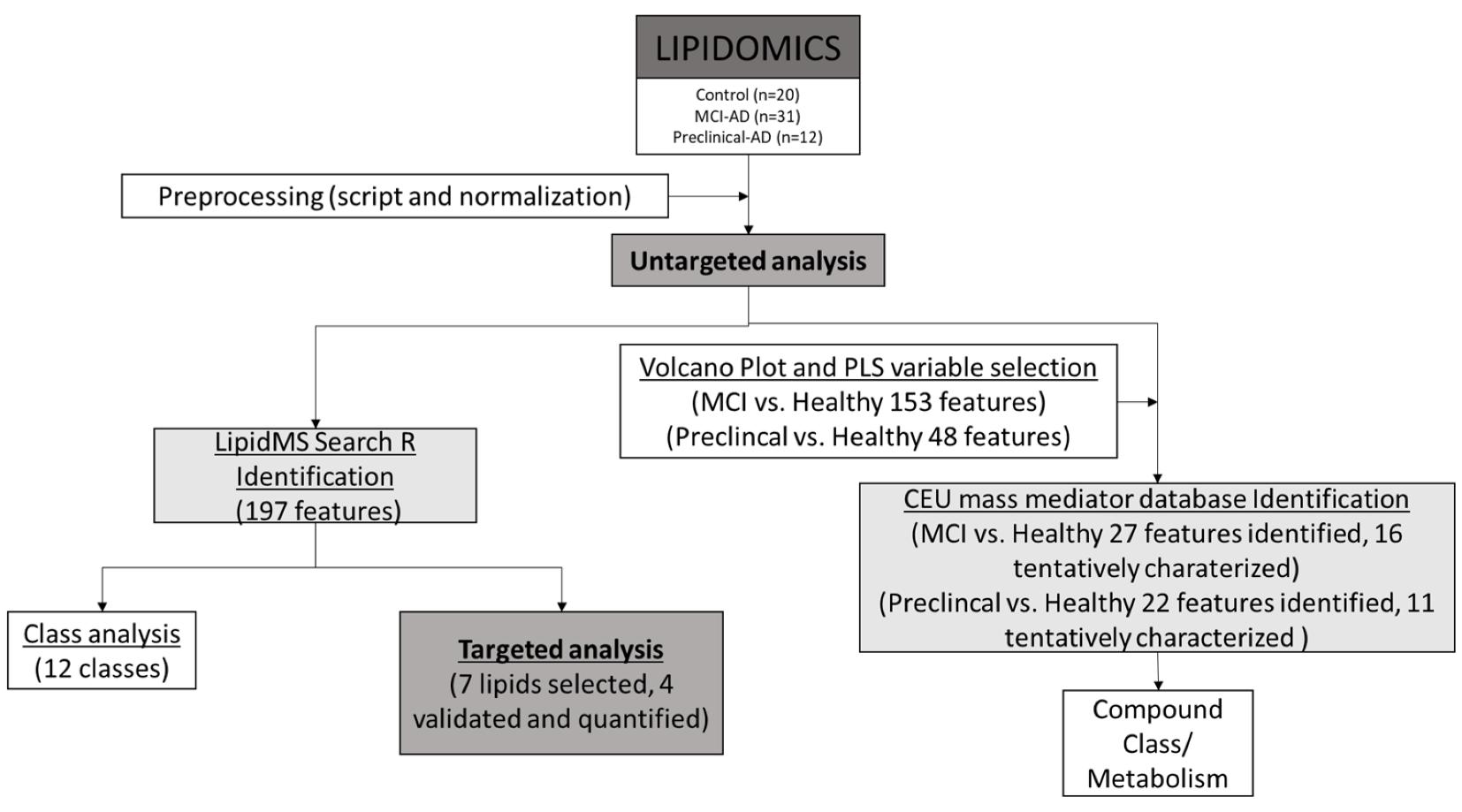
Untargeted plasma screening identifies limited statistically robust features, with high inter-individual variability and unclear lipid-class drivers.
Goal
Increase coverage and confidence in candidate selection prior to committing to targeted validation.
Recommended Path
Discovery (Bundle A) → Expanded statistical and pathway-level refinement
Recommended Services
What You Will Receive
A QC-reviewed differential feature table, multivariate plots (PCA/heatmap), class-level enrichment summaries, and a prioritized lipid shortlist suitable for follow-up validation.
Situation
Candidate lipids identified during discovery do not reproduce across analytical batches, collection waves, or collaborating sites.
Goal
Stabilize quantification and establish a reproducible lipid panel for cohort-level comparison.
Recommended Path
Validation (Bundle B) → Batch-aware statistical reporting
Recommended Services
What You Will Receive
Quantitative LC-MS/MS outputs with internal standard QC review, cross-batch comparability assessment, and a refined marker panel with improved analytical stability.
Situation
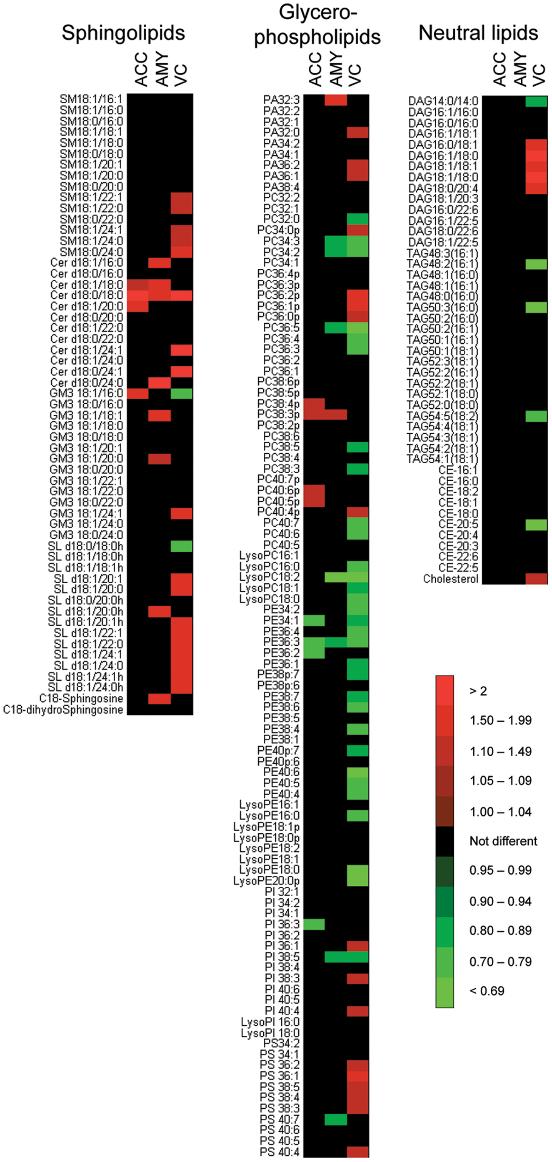
Genetic, transcriptomic, or prior literature evidence implicates sphingolipid or ceramide metabolism in AD/PD biology.
Goal
Quantify species-level sphingolipid changes to support mechanistic interpretation.
Recommended Path
Discovery screening → Targeted confirmation using focused sphingolipid panels
Recommended Services
What You Will Receive
Quantitative sphingolipid and ceramide datasets, species-level concentration tables, and pathway-aligned summaries linking plasma and brain tissue findings.
Situation
Bulk lipidomics indicates significant changes, but regional specificity within brain tissue remains unresolved.
Goal
Determine anatomical distribution of lipid alterations and correlate with defined regions or histological features.
Recommended Path
Targeted validation → Spatial lipid mapping (Deep Insight)
Recommended Services
What You Will Receive
Spatial ion intensity maps, ROI-level comparisons, and region-resolved lipid distribution profiles suitable for integration with histological or molecular datasets.
Situation
Data or literature suggests altered cholesterol metabolism, oxidative stress, or sterol oxidation in AD/PD models.
Goal
Quantify oxysterols and evaluate cholesterol pathway remodeling within plasma or brain tissue.
Recommended Path
Targeted sterol module → Optional spatial confirmation if tissue context is required
Recommended Services
What You Will Receive
Oxysterol quantification tables, cholesterol pathway summaries, and analytically validated sterol profiles supporting neuro-metabolic interpretation.
Situation
Preliminary data indicate bioenergetic stress or membrane remodeling consistent with mitochondrial involvement.
Goal
Differentiate systemic lipid alterations from mitochondria-associated remodeling and prioritize confirmatory markers.
Recommended Path
Discovery profiling → Mitochondria-focused analysis → Targeted confirmation
Recommended Services
What You Will Receive
Mitochondria-enriched lipid profiles, mechanistic pathway interpretation, and a focused marker set for downstream validation.
Situation
Microdissected regions or archival samples restrict available tissue mass.
Goal
Maximize analytical yield while maintaining data robustness and interpretability.
Recommended Path
Fit-for-sample discovery workflow → Focused targeted validation
Recommended Services
What You Will Receive
An optimized analytical plan aligned to input constraints, supported by targeted confirmation to reduce false positives and enhance interpretability.
Situation
Abundance data alone cannot determine whether lipid changes reflect altered synthesis, turnover, or accumulation.
Goal
Introduce metabolic context to support mechanistic conclusions.
Recommended Path
Metabolic flux assessment → Targeted confirmation of key markers
Recommended Services
What You Will Receive
Turnover-oriented analytical outputs that clarify pathway directionality and support mechanistic interpretation in AD/PD neuro-metabolism studies.